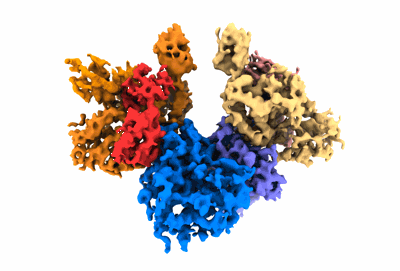
Resolution: 3.06 Å
EM Method: Single-particle
Fitted PDBs: 8bei
Q-score: 0.543
Sendker LF, Lo YK, Heimerl T, Bohn S, Pfister P, Mais N, Sadowska W, Peterson L, Benesch JLP, Marklund E, Bange G, Erb TJ, Schuller JM, Hochberg GKA
To Be Published
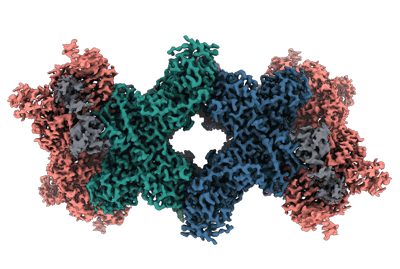
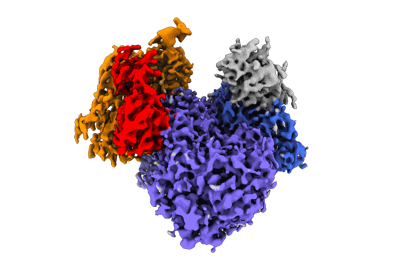
Resolution: 5.91 Å
EM Method: Single-particle
Fitted PDBs: 8rjk
Q-score: 0.277
Sendker FL, Lo YK, Heimerl T, Bohn S, Persson LJ, Mais CN, Sadowska W, Paczia N, Nussbaum E, Del Carmen Sanchez Olmos M, Forchhammer K, Schindler D, Erb TJ, Benesch JLP, Marklund EG, Bange G, Schuller JM, Hochberg GKA
Nature (2024) [ Pubmed: 38600380 DOI: doi:10.1038/s41586-024-07287-2 ]
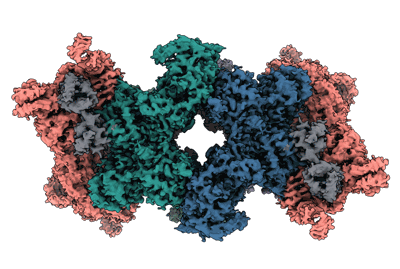
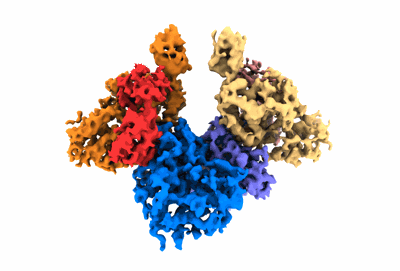
Resolution: 3.34 Å
EM Method: Single-particle
Fitted PDBs: 8rjl
Q-score: 0.504
Sendker FL, Lo YK, Heimerl T, Bohn S, Persson LJ, Mais CN, Sadowska W, Paczia N, Nussbaum E, Del Carmen Sanchez Olmos M, Forchhammer K, Schindler D, Erb TJ, Benesch JLP, Marklund EG, Bange G, Schuller JM, Hochberg GKA
Nature (2024) [ Pubmed: 38600380 DOI: doi:10.1038/s41586-024-07287-2 ]
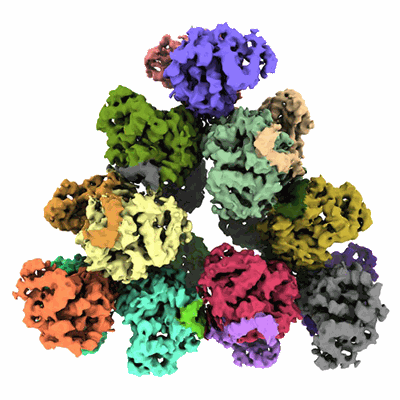
Structure of a first level Sierpinski triangle formed by a citrate synthase
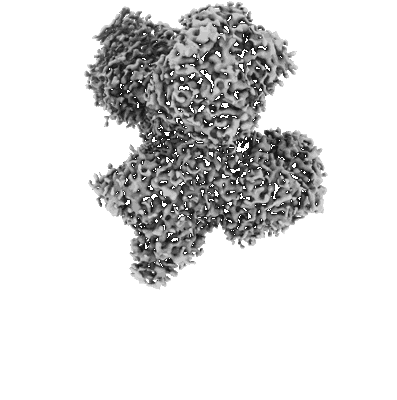
Resolution: 3.93 Å
EM Method: Single-particle
Fitted PDBs: 8an1
Q-score: 0.386
Sendker LF, Lo YK, Heimerl T, Bohn S, Pfister P, Mais N, Sadowska W, Peterson L, Benesch JLP, Marklund E, Bange G, Erb TJ, Schuller JM, Hochberg GKA
To Be Published
Resolution: 3.0 Å
EM Method: Single-particle
Fitted PDBs: 8oh5
Q-score: 0.491
Kumar A, Kremp F, Roth J, Freibert SA, Muller V, Schuller JM
Nat Commun (2023) 14 pp. 5484-5484 [ Pubmed: 37673911 DOI: doi:10.1038/s41467-023-41212-x ]
- Nadph dihydro-nicotinamide-adenine-dinucleotide phosphate (745 Da, Ligand)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Formate dehydrogenase-o, major subunit (126 kDa, Protein from Sporomusa ovata DSM 2662)
- Nad-dependent formate dehydrogenase gamma subunit (18 kDa, Protein from Sporomusa ovata DSM 2662)
- Nad-reducing hydrogenase subunit hoxf (63 kDa, Protein from Sporomusa ovata DSM 2662)
- Cryo-em structure of the electron bifurcating trans-hydrogenase stnabc complex from sporomusa ovata (state 2) (850 kDa, Complex from Sporomusa ovata DSM 2662)
- Nicotinamide-adenine-dinucleotide (663 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
- Flavin mononucleotide (456 Da, Ligand)
Resolution: 3.2 Å
EM Method: Single-particle
Fitted PDBs: 8oh9
Q-score: 0.46
Kumar A, Kremp F, Roth J, Freibert SA, Muller V, Schuller JM
Nat Commun (2023) 14 pp. 5484-5484 [ Pubmed: 37673911 DOI: doi:10.1038/s41467-023-41212-x ]
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Formate dehydrogenase-o, major subunit (126 kDa, Protein from Sporomusa ovata DSM 2662)
- Nad-dependent formate dehydrogenase gamma subunit (18 kDa, Protein from Sporomusa ovata DSM 2662)
- Nad-reducing hydrogenase subunit hoxf (63 kDa, Protein from Sporomusa ovata DSM 2662)
- Zinc ion (65 Da, Ligand)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Cryo-em structure of the electron bifurcating trans-hydrogenase stnabc complex from sporomusa ovata in the oxidised state (850 kDa, Complex from Sporomusa ovata DSM 2662)
- Iron/sulfur cluster (351 Da, Ligand)
CryoEM structure of a tungsten-containing aldehyde oxidoreductase from Aromatoleum aromaticum
Resolution: 3.22 Å
EM Method: Single-particle
Fitted PDBs: 8c0z
Q-score: 0.541
Winiarska A, Ramirez-Amador F, Hege D, Gemmecker Y, Prinz S, Hochberg G, Heider J, Szaleniec M, Schuller JM
Sci Adv (2023) 9 pp. eadg6689-eadg6689 [ Pubmed: 37267359 DOI: doi:10.1126/sciadv.adg6689 ]
- Tungsten-containing aldehyde oxidoreductase from aromatoleum aromaticum (304 kDa, Complex from Aromatoleum aromaticum)
- Aldehyde:ferredoxin oxidoreductase,tungsten-containing (65 kDa, Protein from Aromatoleum aromaticum)
- Similar to ferredoxin:nadh oxidoreductases or nadh oxidases,potential subunit of aldehyde oxidoreductase (45 kDa, Protein from Aromatoleum aromaticum)
- Flavin-adenine dinucleotide (785 Da, Ligand)
- Benzoic acid (122 Da, Ligand)
- Tungsten cofactor (1 kDa, Ligand)
- Iron/sulfur cluster (351 Da, Ligand)
- Iron-sulfur cluster-binding protein potential subunit of aldehyde oxidoreductase (20 kDa, Protein from Aromatoleum aromaticum)
- Magnesium ion (24 Da, Ligand)
Electron bifurcating hydrogenase - HydABC from A. woodii
Resolution: 4.7 Å
EM Method: Single-particle
Fitted PDBs: 7q4v
Q-score: 0.211
Katsyv A, Kumar A, Saura P, Poverlein MC, Freibert SA, T Stripp S, Jain S, Gamiz-Hernandez AP, Kaila VRI, Muller V, Schuller JM
J Am Chem Soc (2023) 145 pp. 5696-5709 [ Pubmed: 36811855 DOI: doi:10.1021/jacs.2c11683 ]
- Iron hydrogenase hydb (64 kDa, Protein from Acetobacterium woodii DSM 1030)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Nicotinamide-adenine-dinucleotide (663 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Homodimer complex of hydabc trimer (300 MDa, Complex from Acetobacterium woodii DSM 1030)
- Iron hydrogenase hyda1 (63 kDa, Protein from Acetobacterium woodii DSM 1030)
- Iron/sulfur cluster (351 Da, Ligand)
- Iron hydrogenase hydc (16 kDa, Protein from Acetobacterium woodii DSM 1030)
- Flavin mononucleotide (456 Da, Ligand)
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 8a5e
Q-score: 0.409
Katsyv A, Kumar A, Saura P, Poverlein MC, Freibert SA, T Stripp S, Jain S, Gamiz-Hernandez AP, Kaila VRI, Muller V, Schuller JM
J Am Chem Soc (2023) 145 pp. 5696-5709 [ Pubmed: 36811855 DOI: doi:10.1021/jacs.2c11683 ]
- Iron hydrogenase hydb (64 kDa, Protein from Acetobacterium woodii DSM 1030)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Electron bifurcating fe-fe hydrogenase hydabc complex from thermoanaerobacter kivui (348 kDa, Complex from Thermoanaerobacter kivui)
- Zinc ion (65 Da, Ligand)
- 2 iron/2 sulfur/5 carbonyl/2 water inorganic cluster (355 Da, Ligand)
- Iron hydrogenase hyda1 (63 kDa, Protein from Acetobacterium woodii DSM 1030)
- Iron/sulfur cluster (351 Da, Ligand)
- Iron hydrogenase hydc (16 kDa, Protein from Acetobacterium woodii DSM 1030)
- 1,4-dihydronicotinamide adenine dinucleotide (665 Da, Ligand)
- Flavin mononucleotide (456 Da, Ligand)
Resolution: 3.78 Å
EM Method: Single-particle
Fitted PDBs: 7q4w
Q-score: 0.253
Katsyv A, Kumar A, Saura P, Poverlein MC, Freibert SA, T Stripp S, Jain S, Gamiz-Hernandez AP, Kaila VRI, Muller V, Schuller JM
J Am Chem Soc (2023) 145 pp. 5696-5709 [ Pubmed: 36811855 DOI: doi:10.1021/jacs.2c11683 ]
- Homodimer complex of hydabc trimer (350 MDa, Complex from Acetobacterium woodii DSM 1030)
- Iron hydrogenase hydb (50 kDa, Protein from Acetobacterium woodii DSM 1030)
- Fe2/s2 (inorganic) cluster (175 Da, Ligand)
- Zinc ion (65 Da, Ligand)
- Iron hydrogenase hyda1 (63 kDa, Protein from Acetobacterium woodii DSM 1030)
- Iron/sulfur cluster (351 Da, Ligand)
- Iron hydrogenase hydc (16 kDa, Protein from Acetobacterium woodii DSM 1030)
- Flavin mononucleotide (456 Da, Ligand)